Can DIY Microscopy Finally Democratize Spatial Biology?
The field of Antibody-Oligonucleotide Conjugate (AOC) techniques continues developing at an impressive pace! Following up our earlier summary of AOC techniques in Scientific Techniques Using Antibody-Oligonucleotide Conjugates, I wanted to highlight a paper that tackles one of the biggest barriers to adoption: cost, accessibility and flexibility.
The Problem
Multiplexed imaging platforms have transformed tissue biology, but commercial systems remain prohibitively expensive and locked to proprietary reagents. This limits access to well-funded core facilities and restricts researchers who need to customize protocols or integrate novel reagents.
The Solution
Fluid-Squid (Simplifying QUantitative Imaging platform Development) combines the open-source Squid microscopy platform with automated fluidics for cyclic imaging, all for around $30k in materials. The compact system has been field-tested from Stanford to Liberia, proving its robustness in diverse settings.
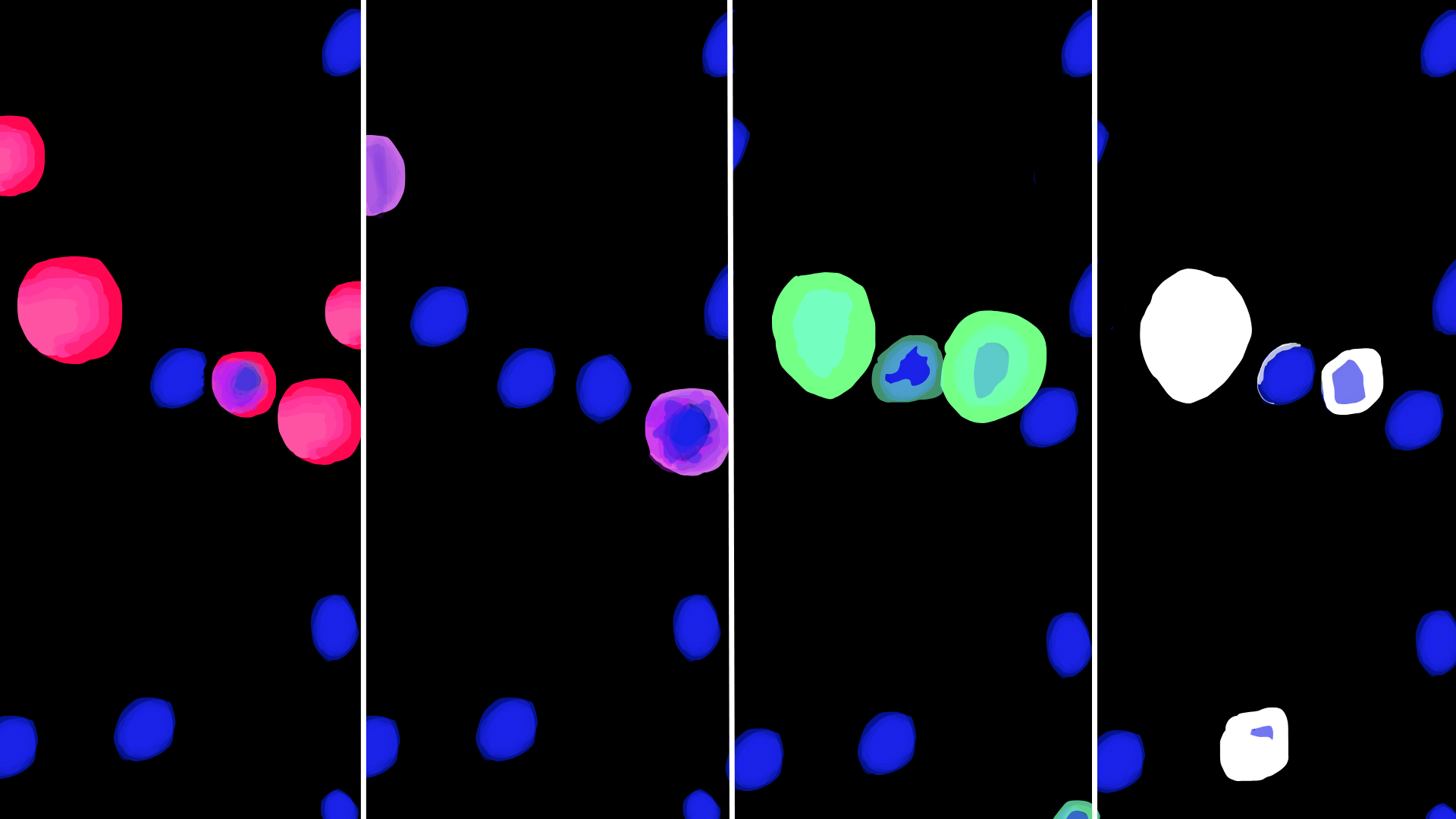
Multiplexed AOC imaging showing distinct cell populations visualized with different fluorescent markers
Key Achievements
- 36-plex AOC imaging of human intestine tissue across a 4.5mm × 4.5mm area at single-cell resolution
- Extended workflows to dissociated single cells (PBMCs and splenocytes) with a 39 AOC panel
- Demonstrated lyophilization of AOC panels, critical for field deployment where cold-chain storage isn't feasible
- Developed AOC conjugation strategy enabling simultaneous staining and rapid titration of 40+ marker panels in single experiments
What Will It Take for Widespread Adoption?
The hardware democratization is impressive, but the real bottleneck may shift to reagents. Labs building custom panels — whether for immuno-oncology or neuroscience ion channel mapping — still need access to AOCs, and the broader availability of ready-to-use AOC reagents could dramatically lower the barrier for prototyping spatial proteomics experiments without being restricted to a specific protocol.