Can Light Replace Wash Steps in DNA-PAINT?
A deeper look at photocaged antibody-oligo conjugates for wash-free multiplexed imaging
A recent paper describes a photocaging-based imaging strategy for multiplexed high-resolution microscopy using DNA labels that can be switched with light. The method, referred to as cagePAINT, is built around a simple but powerful idea: instead of physically exchanging imaging strands between rounds, the binding behaviour of the oligo labels is changed directly with a pulse of violet light.
This matters because multiplexed DNA-PAINT is powerful, but it is not frictionless. In a conventional workflow, each imaging round often requires a fresh buffer exchange to remove one imager strand and introduce the next. That is manageable at low plex, but it becomes increasingly inconvenient as experiments scale. Every wash adds handling time, increases the risk of sample drift, and can compromise overlay accuracy when the goal is nanometre-scale registration across multiple targets. That bottleneck is exactly what this work tries to remove.
Why Sequential DNA-PAINT Still Creates Friction
DNA-PAINT has become one of the most elegant uses of antibody-oligonucleotide conjugates in advanced imaging. Antibodies carry DNA docking strands, and fluorescent imager oligos transiently bind those docking sites to generate localisation events. The approach offers exceptional spatial resolution and, because DNA sequences can be reassigned, strong multiplexing potential. But in practice, that multiplexing usually comes with repeated reagent exchange.
The problem is not just inconvenience. In super-resolution workflows, even subtle perturbations matter. When the specimen is handled repeatedly, there is greater opportunity for mechanical drift, distortion, or slight morphology changes between imaging rounds. That makes the final multi-target reconstruction harder to trust, especially when the biology depends on nanoscale spatial relationships.
The value of this paper is not simply that it adds another DNA-PAINT variant. It addresses one of the format's core operational weaknesses: the need to physically touch the sample every time you want to switch targets.
What the Photocaged Strategy Changes
The paper introduces DNA oligonucleotides carrying photocaging groups that alter whether hybridisation can occur. Illumination with 405 nm light removes or changes those caging effects, allowing target accessibility to be modulated using light rather than liquid handling. In other words, target switching becomes an optical event instead of a wash step.
The design is especially elegant because it uses two complementary behaviours in parallel. One docking strand begins in an active state and can be switched OFF by violet light, while another begins inactive and can be switched ON. Because both targets can be read using the same imager strand sequence, the microscope can move from one target to the next with a brief pulse of light rather than a fresh buffer exchange.
That sounds like a small change, but it is conceptually important. In standard multiplex imaging, the code that determines which target is visible is carried by reagent exchange. Here, that code is shifted into the oligo design itself. The AOC becomes not just a recognition reagent, but a light-responsive logic element inside the assay.
Key insight: The AOC becomes not just a recognition reagent, but a light-responsive logic element inside the assay — target switching becomes an optical event instead of a wash step.
How the Antibody-Oligo Conjugates Were Engineered
From an AbOliGo perspective, the most interesting part of the paper is the oligo chemistry. The team synthesised a Methyl-DEACM caged thymidine phosphoramidite and incorporated it into DNA docking strands by solid-phase oligonucleotide synthesis. These coumarin-derived photocages interfere with hybridisation until the strand is exposed to the appropriate light input.
That means the crucial functional change does not come from the antibody itself. The antibody still provides target specificity. The programmable behaviour sits in the oligonucleotide cargo attached to the antibody. Once those modified docking strands are coupled to an antibody, the result is effectively a light-switchable AOC: the same biological binder, but with target visibility controlled optically rather than purely chemically.
This is a useful reminder that in imaging assays, the oligo portion of an AOC is not passive. Sequence, accessibility, secondary structure, steric bulk, and in this case photocaging chemistry all influence whether the final conjugate behaves as intended. That same general principle sits behind many of the methods covered across the AbOliGo knowledge hub, from proximity ligation to cyclic imaging and DNA-PAINT.
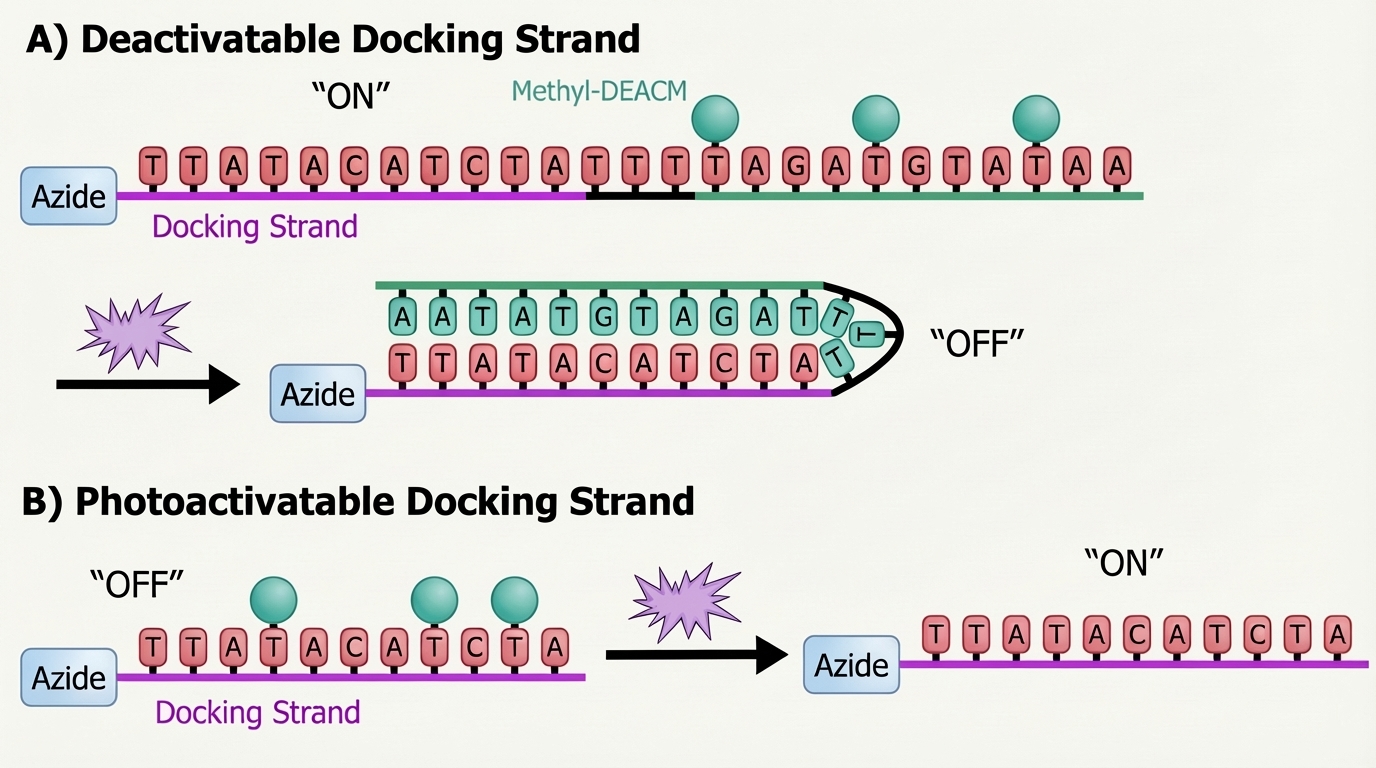
Design of the photodeactivatable and photoactivatable docking strands A and B. Initially, A is accessible for hybridisation to the complementary imager strand, while B remains in its "OFF" state. Upon 405 nm (purple star) illumination, the Methyl-DEACM cage groups are removed: oligo A adopts a closed, inactive ("OFF") conformation, whereas B becomes activated and available to bind the complementary imager strand.
What Was Demonstrated
The method was demonstrated across confocal microscopy, DNA-PAINT single-molecule localisation microscopy, and STED microscopy, which is important because it shows the concept is not tied to a single imaging mode. The study also describes two-target confocal imaging of microtubules and mitochondria, illustrating practical target switching within the same specimen.
That breadth gives the paper extra weight. It suggests the photocaging concept is a general imaging control strategy rather than a narrow one-off trick. For the field, that is probably the real advance: not just a new sequence set, but a new way of thinking about how and when an AOC should be visible during an experiment.
Why This Matters Beyond One Super-Resolution Paper
The immediate application is clear: less buffer exchange, less handling, and potentially better registration fidelity in multiplexed imaging. But the broader relevance may be even more interesting. If antibody-bound oligos can be turned on and off cleanly with light, then signal generation no longer has to depend only on reagent addition, washing, or irreversible amplification steps.
That opens up a bigger design space. A light-switchable oligo layer could, in principle, be useful anywhere researchers want tighter spatial or temporal control over when a target becomes readable. Potential future directions include controlled sequential readout in dense tissues, reduced perturbation in delicate samples, optical gating of proximity-style workflows, or more automated multiplexed imaging systems where fluidics are simplified and target switching is handled in software and illumination.
Key takeaway: The future of AOCs is not only about attaching DNA to antibodies. It is about engineering what that DNA can do once it gets there.